
A comprehensive multi platform software for microarray data analysis
Genowiz™ is a powerful gene expression analysis program that has been designed to store, process and visualize gene expression data efficiently. It includes a suite of advanced analysis methods from which researchers can select appropriate analysis methods for their dataset.
Genowiz™ allows researchers to organize experimental information (MIAME), import data files quickly and easily, work simultaneously with multiple experiments, import gene annotation files, pre-process and normalize data, perform cluster analysis, classify and view gene information, perform functional classification and track down intricate correlations in data by performing pathway analysis. All analysis performed is saved in database and can be retrieved as required.
OVERVIEW
Ten great reasons to try Genowiz™:
Proven and Appreciated by Life Scientists:
Genowiz™ has been used to study plant. Prokaryotic, Eukaryotic and clinically relevant microarray expression data.
Easy to learn:
User-friendly organization of features, GUIs and visualization tools which provide knowledge rather information.
Validated Results:
Accurate expression results validated against gold standards like R-package.
Microarray platform independent:
Supports all proprietary and open data formats e.g. Affymetrix, Aglilent. Also upload unique formats using customized uploader.
Comprehensive Statistical and Bio-Analysis:
All major Normalization , Clustering, Classification well integrated with GO, Pathways and Gene Lists comparisons.
Save time on analysis:
Using built-in or user designed AutoGuide workflow module.
Affordable Price:
ust pay once! Perpetual Licensing.
24 X 5 Free Support:
Free Live web demonstration and post sale e-mail support for the life.
Consultative approach of Development:
We continuously improve the software based on the latest scientific developments and your feedback.
Upgrade Price:
First year upgrade free. Minimal Upgrade fee thereafter and Dedicated support contract(Optional).
FEATURES
Data and Gene List Import
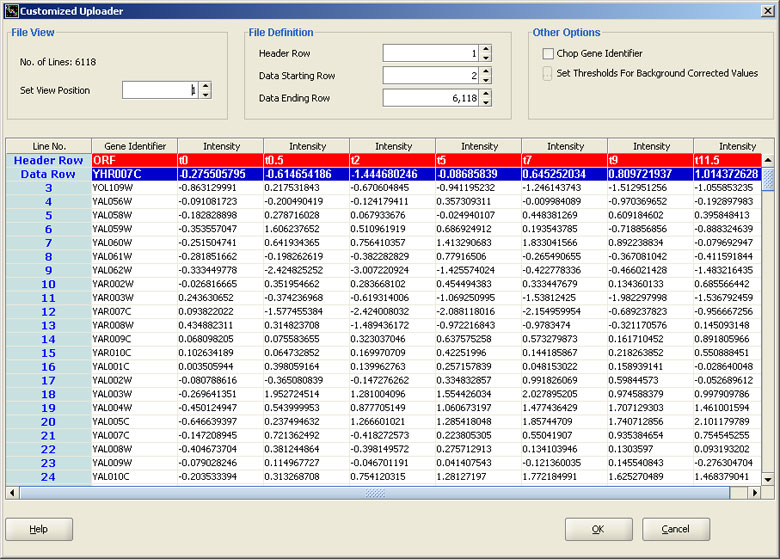
Genowiz™ supports a wide range of cDNA and Affymetrix data formats. Users can directly import .CEL and .CHP files into Genowiz™. Users can upload data in customized formats also. Customized uploader allows users to add and save new data formats. Automatic Uploader can then identify these formats. At the time of data import replicate genes are filtered, dye swap samples and thresholds for maximum and minimum values can be specified. PLIER, RMA and MAS 5.0 algorithms for Affymetrix probe level normalization and summarization are present.
MIAME
Minimum Information About a Microarray Experiment (MIAME) facilitates adoption of standards for microarray experiment annotation and data representation. Genowiz™ focuses on establishing standard microarray experimental data repositories and information sharing within the scientific community. Researchers can also exchange MIAME data by using MAGE ML document exchange format.
Data Transformation, Normalization and Filtrationt
In any type of expression analysis, pre-processing of data to reduce undesirable variation among datasets and to bring data to a common platform is a vital step. Genowiz™ provides users with a wide range of data transformation, normalization and filtration tools. These include:
- Data transformation options such as imputation of missing values, log transformations, mean/median, Z-transformation, subtract control, divide by control, and scaling.
- Normalization techniques such as normalization for dye swap replicates, cDNA raw data normalization options (cDNA Loess and Print tip Loess) and quantile normalization. Separate normalization techniques are provided for cDNA and Affymetrix arrays. Normalization can be done using all genes or control genes.
- Filter data based on replicate samples, fold change, mean, standard deviation, calls and missing values. Replicate samples are handled using various parametric/non-parametric tests like Two Way ANOVA, Mann Whitney U Test, Kruskal Wallis Test, and One Sample t-Test. Multiple testing correction options like Bonferroni/FDR can be applied to reduce false positives.
Data Analysis and Visualization
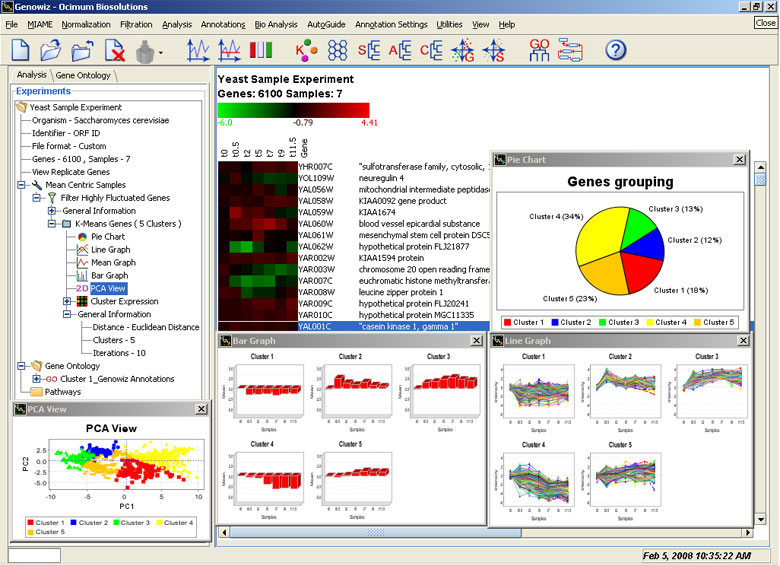
Genowiz™ is equipped with several data analysis tools. Complete with informative graphics, it is an excellent tool for interpretation of biologically meaningful results. Some of these tools include partition clustering, hierarchical clustering, SOM, PCA, gene shaving and discriminant PCA and SVM. Option for merging clusters of interest has also been provided.
Partition Clustering (K-Means)
This tool classifies genes or samples in user-defined groups using distance parameters. The obtained clusters can be re-clustered. Re-clustering utility helps scientists pick a set of genes of their interest. A 2D PCA view shows the distribution of genes in various clusters.
Hierarchical Clustering
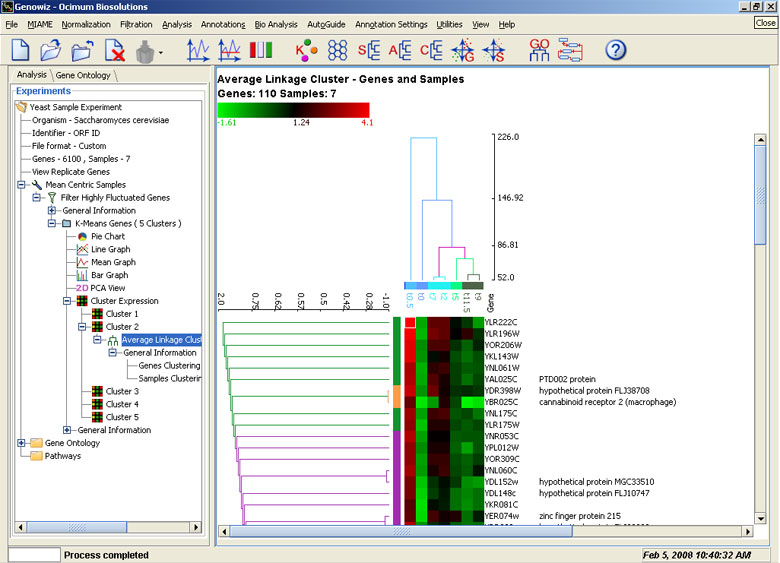
One of the most important tools for studying relations between genes, this tool creates a dendrogram based on the relative distance between genes. The different optional parameters help the user in correctly determining the relationship between two genes. Models of analysis include single linkage, complete linkage and average linkage clustering. Genes, samples, or both together can be clustered.
Self Organizing Maps
two-way classification of genes into clusters based on novel artificial neural networks is an integral feature of data clustering tools in Genowiz™. This gives a deeper insight into clusters, as neighboring clusters are very similar to each other.
Principal Component Analysis
This tool involves a mathematical procedure that transforms a number of (possibly) correlated variables into a (smaller) number of uncorrelated variables called principal components. These provide an insight into existent variability in the data.
Gene Shaving
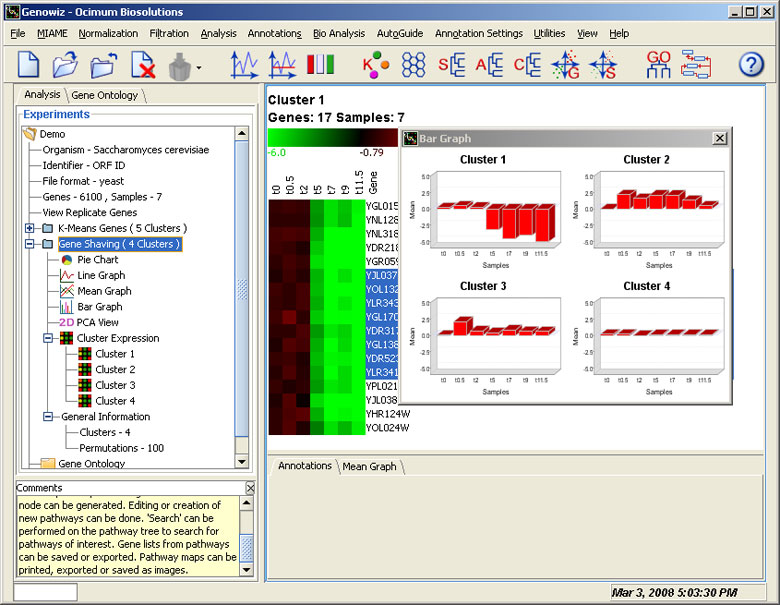
This method identifies subsets of genes with coherent expression patterns and large variation across conditions. Gene shaving differs from hierarchical clustering and other methods of gene expression analysis in that genes may belong to more than one cluster.
Classification
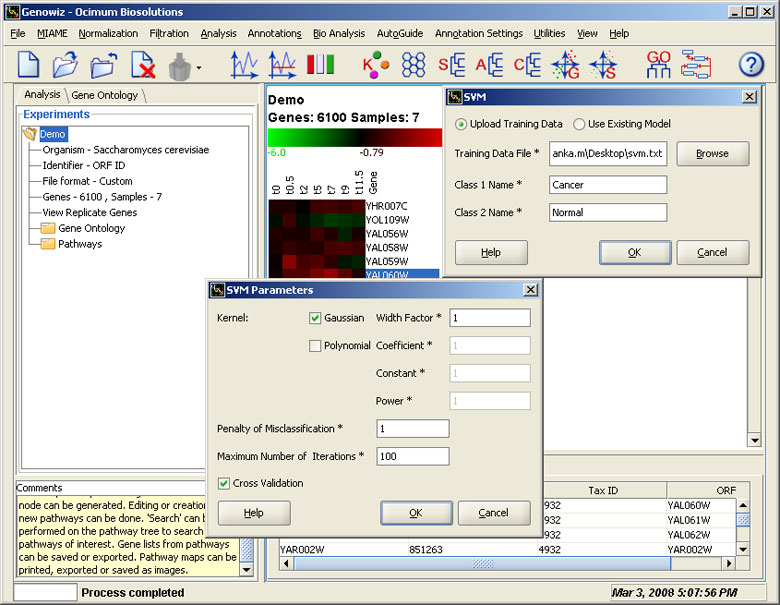
Classification algorithms are used to classify samples, based on information from similar samples with known classes that are available in training data. In Genowiz™, Support Vector Machines (SVM) and Discriminant PCA are used to predict classes for unclassified samples.
Box Plot & Scatter Plot
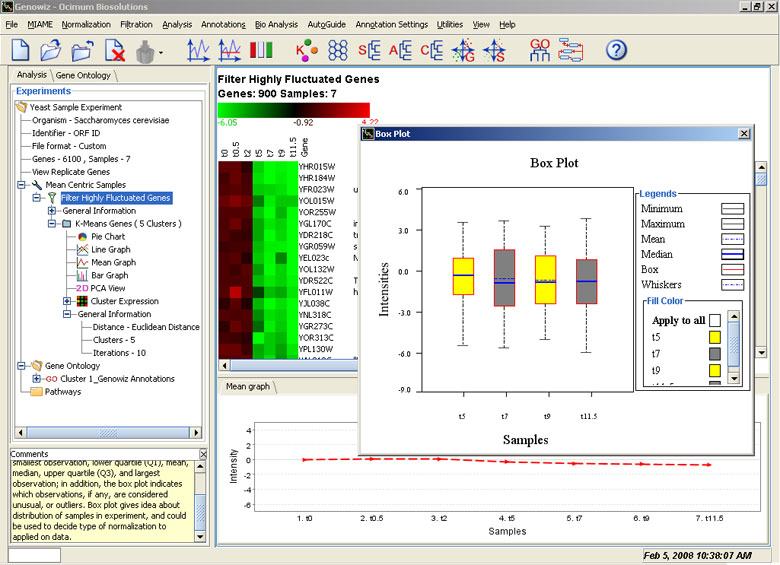
Sample distribution can be visualized and compared within an experiment using Box plots and Scatter plots. Fold change lines can be drawn on scatter plots. Up and down regulated genes can be color coded and selected genes can be saved as a node. Additional utility options have also been provided.
Biological Analysis
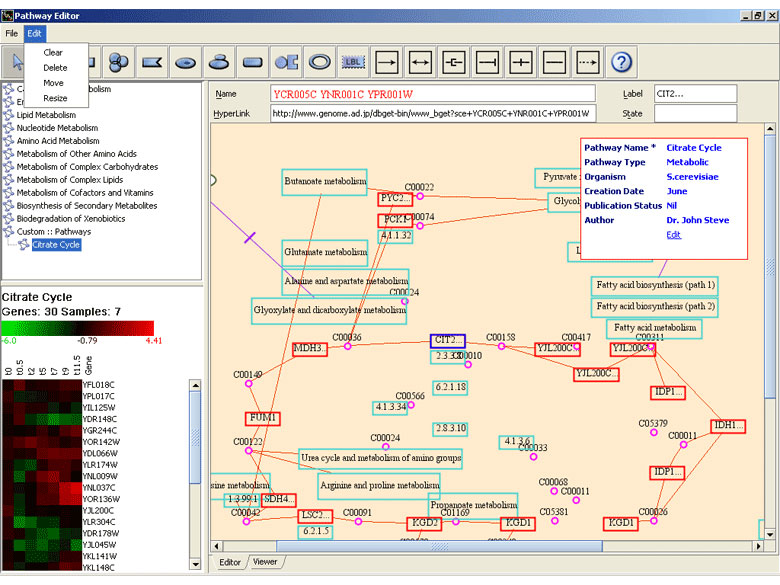
Genowiz™ annotates genes and classifies them into functional categories (Gene Ontology). Option for importing annotation files and NetAffx™ chips information is also provided. Integrated pathways module aids researchers in understanding pathways in relation to expression data. Pathway maps edited/created can be associated with author details too. Coupled with biological information, gene ontology and pathway classification, it forms an excellent tool in understanding biological systems. Annotations tab displays selected gene(s) annotations. Search can be performed on gene ontology and pathway tree to find ontologies or pathways of interest.
Saving Gene Ontology and Pathway Analysis
Gene ontology and pathway analyses performed on different experiments are saved to the database. Functional clusters of interest can be saved as gene lists and can be compared using Gene List Comparison feature.
Add Organisms
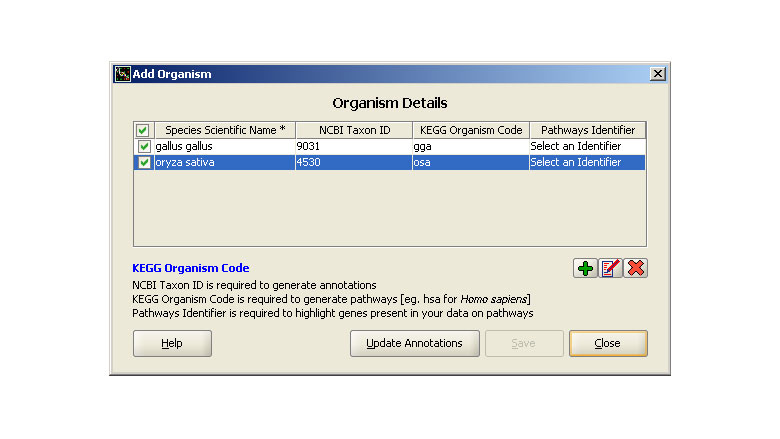
Genowiz™ supports seven organisms by default. Researchers can add organisms and genomes of interest and update corresponding annotations.
Automatically update Genowiz™ Annotations
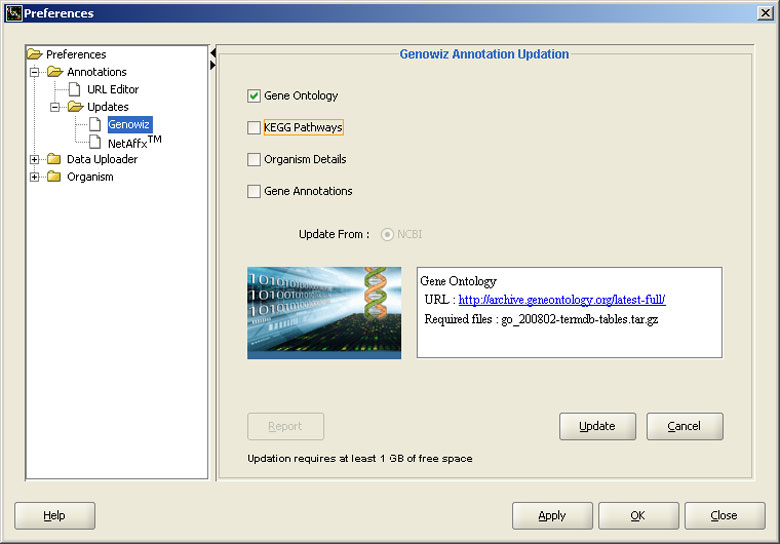
Annotations can be updated for organism(s), either by connecting to respective websites or from local disk (in case of internet connection, firewall, licensing or download problems). Information regarding gene ontology, pathways, organisms and annotations from Entrez Gene database can be updated.
Utilities
Few of the utility options that add value to the analyses performed:
- Gene List Comparison: Subtle relations among datasets can be probed using this feature. Up to four gene lists can be compared with a Venn diagram representation.
- Pattern Simulation: For an expression pattern defined, all genes with a similar expression pattern can be listed. This gene list can be saved and exported.
- Gene Tracking: Important genes or genes of interest can be tagged and tracked throughout the analysis.
Import/Export Experiments
All analyses performed on an experiment can be exported to specified location and same can be imported into Genowiz. This facilitates sharing of experiments.
Work Flows
Automated work flows for single, double and dye swap experiments effectively guide researchers in their analysis. User created work flows can be saved and applied to experiments.
Technical Support
Ocimum’s technical support staff is available for 24 hours, five days a week to answer your questions about Genowiz™ over phone, e-mail and web chat. Previously answered questions by the support staff are available on the website. A Quick Start Guide provided along with the application, assists users to work with various modules. The Technical Support feature in Genowiz (Help > Technical Support) enables users to send a mail to support team for further assistance.
